Physical Address
304 North Cardinal St.
Dorchester Center, MA 02124
Physical Address
304 North Cardinal St.
Dorchester Center, MA 02124

There are 2,236 active molecular biology databases in the world as of January 2025, according to the Nucleic Acids Research Molecular Biology Database Collection. Each one uses its own schema, identifiers, API conventions, rate limits, and licensing terms. The number of unified, agent-ready APIs that allow a developer or an AI agent to query across these sources through a single interface? Zero.
This is the central finding of a strategic feasibility study on the viability of building biodata infrastructure for AI agents, a normalisation and interoperability layer that sits between fragmented biology databases and the AI systems that need to consume them. The question was whether this gap represents a real, commercially viable opportunity. The answer is yes but with important caveats.
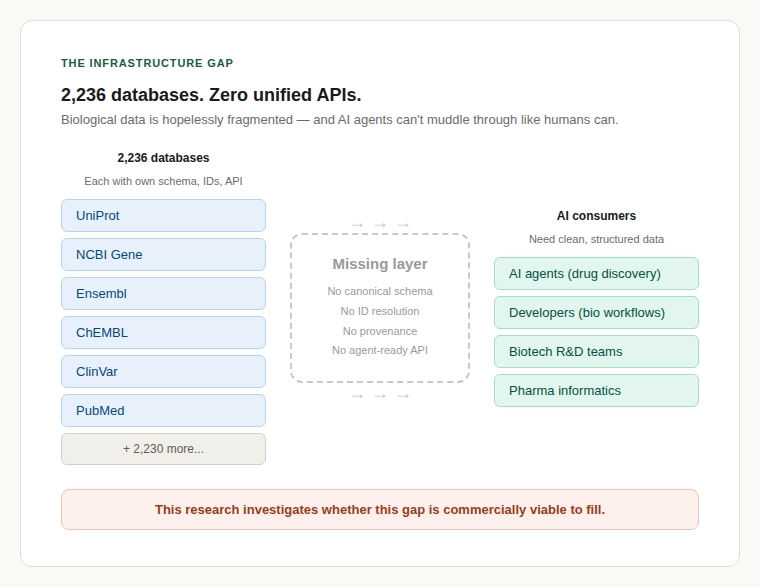
The fragmentation of bioinformatics data is one of the oldest complaints in computational biology. But it has historically been treated as an inconvenience rather than a commercial bottleneck. Two things have changed.
The scale of the problem has grown beyond what manual solutions can handle. The NAR database collection grew from 1,170 resources in 2016 to 2,236 in 2025 nearly doubling in under a decade. The 2026 issue added another 84 new databases, with the collection settling at 2,173 after pruning dead URLs. Each new database adds another node to the integration problem. The combinatorial complexity of linking data across sources grows faster than the number of sources themselves.
AI agents have entered the picture and they can’t muddle through. Unlike human researchers who apply domain intuition to inconsistent data, an AI agent querying UniProt, then NCBI Gene, then ChEMBL, then ClinVar needs structured, consistent, machine-readable responses. It gets none of those things today. This is the core tension: the fastest-growing class of biodata consumers is the least equipped to handle the current infrastructure.
| Metric | Value | Source |
|---|---|---|
| Active bioinformatics databases | 2,236 (Jan 2025) | NAR Molecular Biology Database Collection |
| Published biological AI agent systems | 115 (Jan 2023 – Sep 2025) | Briefings in Bioinformatics (2026) |
| AI drug discovery companies worldwide | 530+ (Oct 2025) | FounderNest Pharma Investment Trends |
A peer-reviewed survey published in Briefings in Bioinformatics in early 2026 conducted a structured literature review of biological AI agent systems published between January 2023 and September 2025. The authors searched arXiv, PubMed, bioRxiv, and Google Scholar. They applied strict inclusion criteria: a system had to demonstrate multi-step reasoning, autonomous task planning, persistent state or memory management, and closed-loop interaction with biological tools or databases. Single-turn chatbots and one-shot LLM wrappers were excluded.
They found 115 representative studies meeting these criteria.
Before January 2023, this number was effectively zero. The entire field of biological AI agents, not AI applied to biology generally, but agentic systems that plan, reason, use tools, and operate autonomously on biodata, materialised in under three years.
The current count of biological AI agents is small. The growth trajectory is what matters. From zero to 115 published systems in under three years represents one of the fastest-emerging demand signals for a new category of biodata infrastructure.
The commercial layer is growing in parallel. Over 530 companies worldwide now focus on AI-powered drug discovery, according to FounderNest’s pharma investment tracking. The TechBio sector raised $5.3 billion between Q3 2023 and Q2 2024. Xaira Therapeutics launched with over $1 billion in seed funding. Recursion Pharmaceuticals acquired Exscientia for $688 million. Isomorphic Labs closed a £449 million Series A.
Real capital is flowing into AI systems that depend on biological data. And those systems are running headfirst into the infrastructure gap.
It is easy to say “bioinformatics data is fragmented” without appreciating the operational reality. Here is what it means for someone building a working system.
Identifier mismatch (the hardest unsolved problem). The same human gene can be referred to by an Ensembl ID (ENSG00000141510), an NCBI Gene ID (7157), an HGNC symbol (TP53), a UniProt accession (P04637), and a RefSeq ID (NM_000546). Mapping between these is version-dependent, lossy at the edges, and breaks differently across organisms. Every team that works across databases builds their own identifier resolution pipeline. Every one of them discovers the same edge cases the hard way.
Schema inconsistency across every database. A protein record in UniProt has completely different fields and structure than one in PDB, NCBI Protein, or the AlphaFold Database. There is no shared data model. Every database returns data in its own format for example, JSON here, XML there, TSV somewhere else. Building an ETL pipeline for even 10 bioinformatics sources consumes months of engineering time.
Rate limits that break AI agent workflows. NCBI’s E-utilities API enforces a rate limit of 3 requests per second without an API key, 10 with one. An AI agent conducting a multi-step drug-target analysis might need dozens of sequential queries. At 3 per second, even simple agentic workflows become bottlenecked.
Provenance loss across database aggregation. When data flows through multiple databases, the original evidence basis often disappears. A drug-target association might be backed by a single in-vitro experiment, a large clinical trial, or a computational prediction. When aggregated downstream, the distinction is frequently lost.
Licensing ambiguity for commercial use. Public bioinformatics databases are not uniformly “open.” UniProt uses CC BY 4.0 (permissive). ChEMBL uses CC BY-SA 3.0 (share-alike). KEGG restricts commercial use. DrugBank requires a commercial license. The patchwork creates legal risk for any company building biodata infrastructure that aggregates or re-serves this data.
| Technical Dimension | Severity (1–5) | Assessment |
|---|---|---|
| Identifier mismatch | 5/5 | Same entity has different IDs across every database. Mapping is version-dependent and lossy. |
| Cross-database joining | 5/5 | Linking gene → protein → drug → clinical evidence requires bespoke pipelines for each path. |
| Schema inconsistency | 4/5 | Every source uses its own data model. No shared canonical schema exists. |
| Ontology mismatch | 4/5 | GO, MeSH, SNOMED, ChEBI different classification systems that don’t neatly align. |
| Provenance visibility | 4/5 | Evidence basis (experiment, prediction, curation) often lost in aggregation. |
| Rate limits | 4/5 | NCBI: 3–10 req/s. Hard bottleneck for any agent-scale workload. |
| Licensing complexity | 4/5 | Patchwork of CC-BY, CC-BY-SA, proprietary terms creates commercial risk. |
| API quality | 3/5 | Ranges from excellent REST (UniProt) to FTP-only dumps (DrugBank). |
| Update frequency | 3/5 | Weekly to quarterly updates require constant sync monitoring. |
| Access restrictions | 3/5 | Most public databases are open; exceptions (KEGG, DrugBank) are significant. |
Given how obvious and long-standing this problem is, you would expect someone to have built the solution. They have not.
BioThings API (Scripps Research) is the closest thing to a unified biodata API. It powers MyGene.info, MyChem.info, and MyVariant.info. But it is an academic project with no commercial support, no canonical schema across sources, and no optimisation for AI agent consumption.
Open Targets Platform integrates genetics, genomics, drugs, and disease data into one excellent resource. But it is designed as a web application for human researchers, not as developer API infrastructure for AI agents.
MCP server builders scattered open-source projects that expose individual databases (PubMed, UniProt) to LLM agents via Model Context Protocol. Each covers one database, with no normalisation, no cross-source identifier resolution, and no provenance tracking.
Nobody has built a commercial product that takes data from multiple biology databases, normalises it into a canonical schema, resolves identifiers across sources, tags provenance, and delivers it through a unified API and MCP interface that AI agents can consume directly.
Where the whitespace is: The moat is not access but it is harmonisation. Anyone can proxy an API call. Solving cross-database identifier resolution and delivering clean, joined, provenance-tracked biodata is genuinely hard engineering that creates a defensible position.
The global bioinformatics market is valued at approximately $14–20 billion in 2024, growing at 13–19% CAGR, projected to reach $50–120 billion by the early 2030s, according to a consensus of reports from SkyQuest, Fact.MR, Fortune Business Insights, DataBridge, Mordor Intelligence, and Research and Markets. The narrower AI-in-drug-discovery sub-market is estimated at $2.4–4.6 billion, growing at 24–30% CAGR.
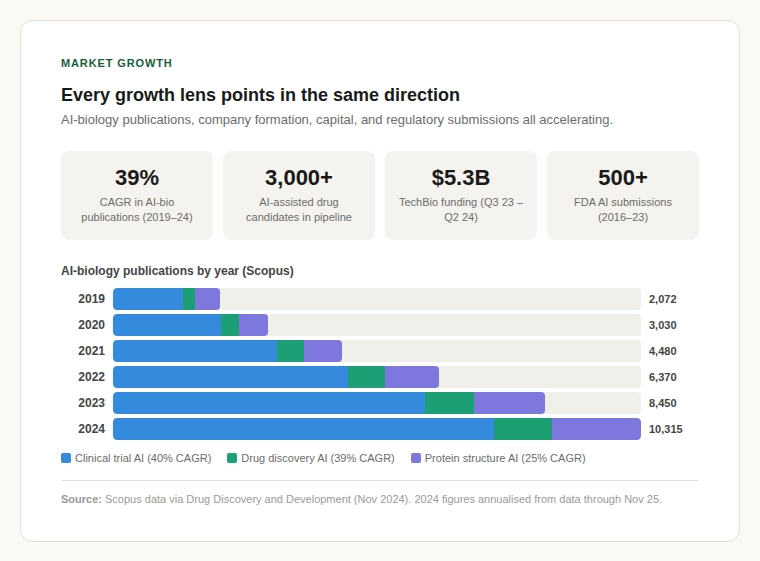
Scopus publication data shows AI-biology publications grew from roughly 2,000 in 2019 to over 10,300 in 2024 which is a sustained 39% CAGR over five consecutive years (Drug Discovery and Development, Nov 2024). The pharmaceutical pipeline now features over 3,000 AI-assisted drug candidates (GlobalData), and more than 500 FDA submissions have included AI components between 2016 and 2023.
The bioinformatics market is valued at $14–20 billion and growing at 13–19% CAGR. Every growth lens including publications, company formation, capital deployment, regulatory submissions points in the same direction: accelerating demand for better biodata infrastructure.
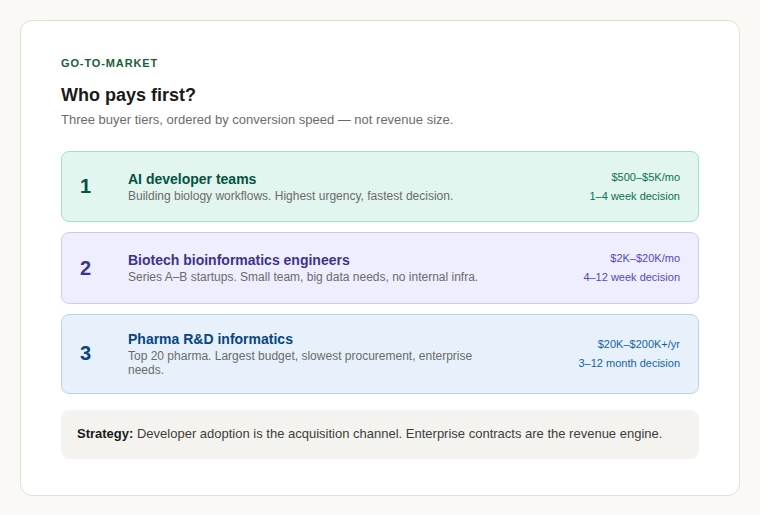
A common failure mode for infrastructure startups is building something technically impressive that nobody writes a cheque for. The buyer landscape breaks into three tiers.
Tier 1: AI developer teams building biology workflows. Highest urgency, lowest tolerance for manual ETL, most natural fit for API and MCP-native delivery.
Tier 2: Bioinformatics engineers at Series A–B biotech startups. Small team, big data needs, no capacity for internal data infrastructure.
Tier 3: Pharma R&D informatics at top-20 companies. Slow procurement and enterprise-grade requirements. Medium-term target, not launch target.
The strategic logic: developer adoption is the acquisition channel; enterprise contracts are the revenue engine.
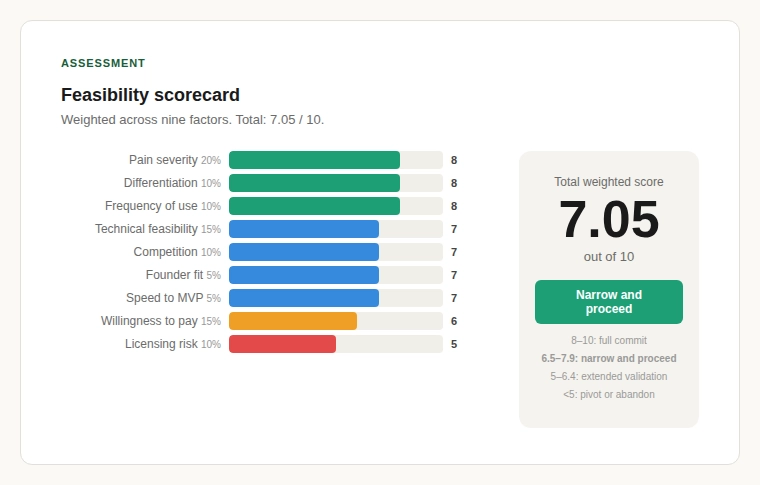
A weighted scorecard across nine factors of pain severity, frequency of use, willingness to pay, technical feasibility, licensing risk, competition intensity, differentiation potential, founder fit, and speed to MVP produces the following result.
| Factor | Weight | Score | Weighted |
|---|---|---|---|
| Pain severity | 20% | 8 | 1.60 |
| Frequency of use | 10% | 8 | 0.80 |
| Willingness to pay | 15% | 6 | 0.90 |
| Technical feasibility | 15% | 7 | 1.05 |
| Data rights / licensing | 10% | 5 | 0.50 |
| Competition intensity | 10% | 7 | 0.70 |
| Differentiation potential | 10% | 8 | 0.80 |
| Founder fit | 5% | 7 | 0.35 |
| Speed to MVP | 5% | 7 | 0.35 |
| Total | 100% | 7.05 / 10 |
A score of 7.05 places this in the “narrow and proceed” zone: above 6.5 (worth pursuing) but below 8.0 (full conviction). The opportunity is real but requires disciplined focus, a tight initial wedge, not a platform play on day one.
The weakest score is licensing risk (5/10). The strongest are pain severity (8/10), differentiation potential (8/10), and frequency of use (8/10). The problem is acute, the competitive whitespace in biodata infrastructure is real, and the usage pattern is daily and recurring.
Verdict: Narrow and proceed. Build a developer-first, API-native biodata product covering 10–15 of the most-used bioinformatics sources in drug-target research. Nail canonical schema quality and identifier resolution before adding breadth. Deliver through REST and MCP. Target AI agent builders first. Plan the enterprise upsell at 12–18 months.
⚠️ Reasons for caution: The buyer is diffuse, identifying who writes the cheque early is critical. Licensing complexity across bioinformatics databases must be addressed at the architecture level. And there is always the risk that large platform players (AWS, Google Cloud Life Sciences, Illumina) build adjacent products. The moat must be built through depth of data harmonisation, not breadth of access.
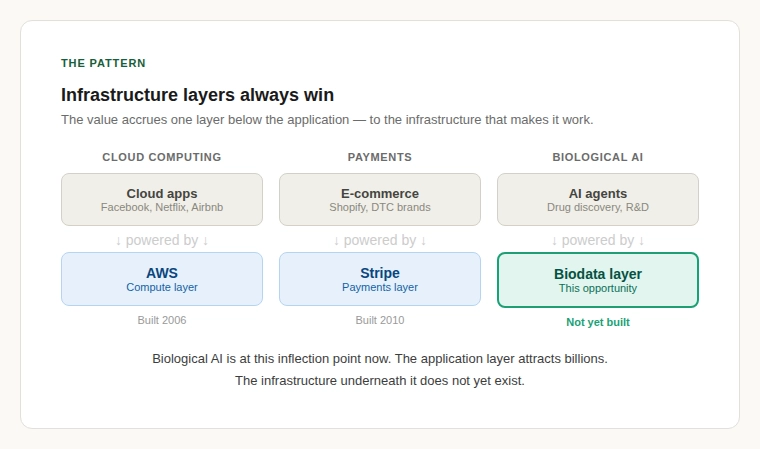
There is a pattern in technology markets. When a new class of application emerges, mobile apps, cloud microservices, AI agents, the initial excitement focuses on the application layer. But the real value often accrues one layer down: to the infrastructure that makes the application layer work. AWS did not build Facebook. It built the cloud infrastructure layer that made thousands of internet startups easier to launch and scale. Stripe did not build e-commerce; it built the payments infrastructure that made e-commerce frictionless.
The biological AI agent space is at this inflection point now. The application layer including the autonomous labs, the AI scientists, the agentic drug discovery platforms is attracting billions in capital. But underneath it all, every one of these systems faces the same problem: getting clean, reliable, cross-referenced biological data into formats that machines can actually use.
Based on this research, I believe the opportunity to build biodata infrastructure for AI agents is genuine. The fragmentation across 2,200+ bioinformatics databases is verified. The demand from 115+ biological AI agents is growing exponentially. The competitive whitespace is clear. And the timing with the emergence of MCP, tool-use frameworks, and the broader agentic AI wave is structurally favourable.
It is not a slam dunk. A feasibility score of 7.05 does not mean guaranteed success. It means the preconditions are met and the risks are identifiable and manageable.